|
If you cannot see the track, check it is enabled under the Annotations and Tracks tab to the right of the viewer. Once the file is imported, the annotations will be displayed as a track on your sequence file. If you use the wrong reference sequence the annotations may be in the wrong location. For example, if you have a GFF file for the human genome build GRCh37, then you must use this version and not GRCh38. You must also ensure that you import the file onto the same sequence that was used to generate the annotations. Note that it is critical that the sequence names match the sequence IDs in the annotation file, otherwise Geneious will not be able to determine which sequence to import the annotations onto. 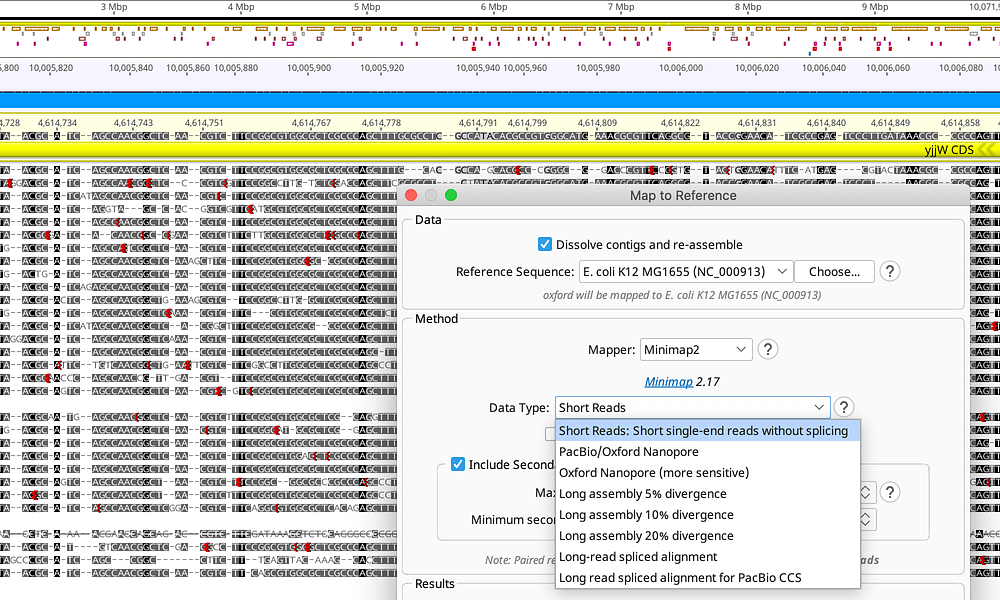
Trim with BBDuk Select the paired list, go to menu Annotate & Predict Trim using BBDuk. Select this file, and you’ll see that each sequence is tagged with a forward or reverse tag. If Geneious cannot find a matching sequence it will import the annotations onto a blank sequence with ? for all bases. In this tutorial, this step has already been done and you are provided with one file of paired reads, with an insert size of 500 bp. If you have the sequence in fasta format and Geneious has not already offered to import that file, choose Import the sequence from a file. If you have not selected the sequence(s), but are in the folder containing the sequence(s), choose Find a sequence with the same name in this Geneious folder. If you already have the sequence(s) in Geneious and have selected the sequence documents prior to importing the annotation file, choose Use one of the selected sequences. If Geneious does not find a matching fasta file in the folder, it will ask where you want to get the sequence from: 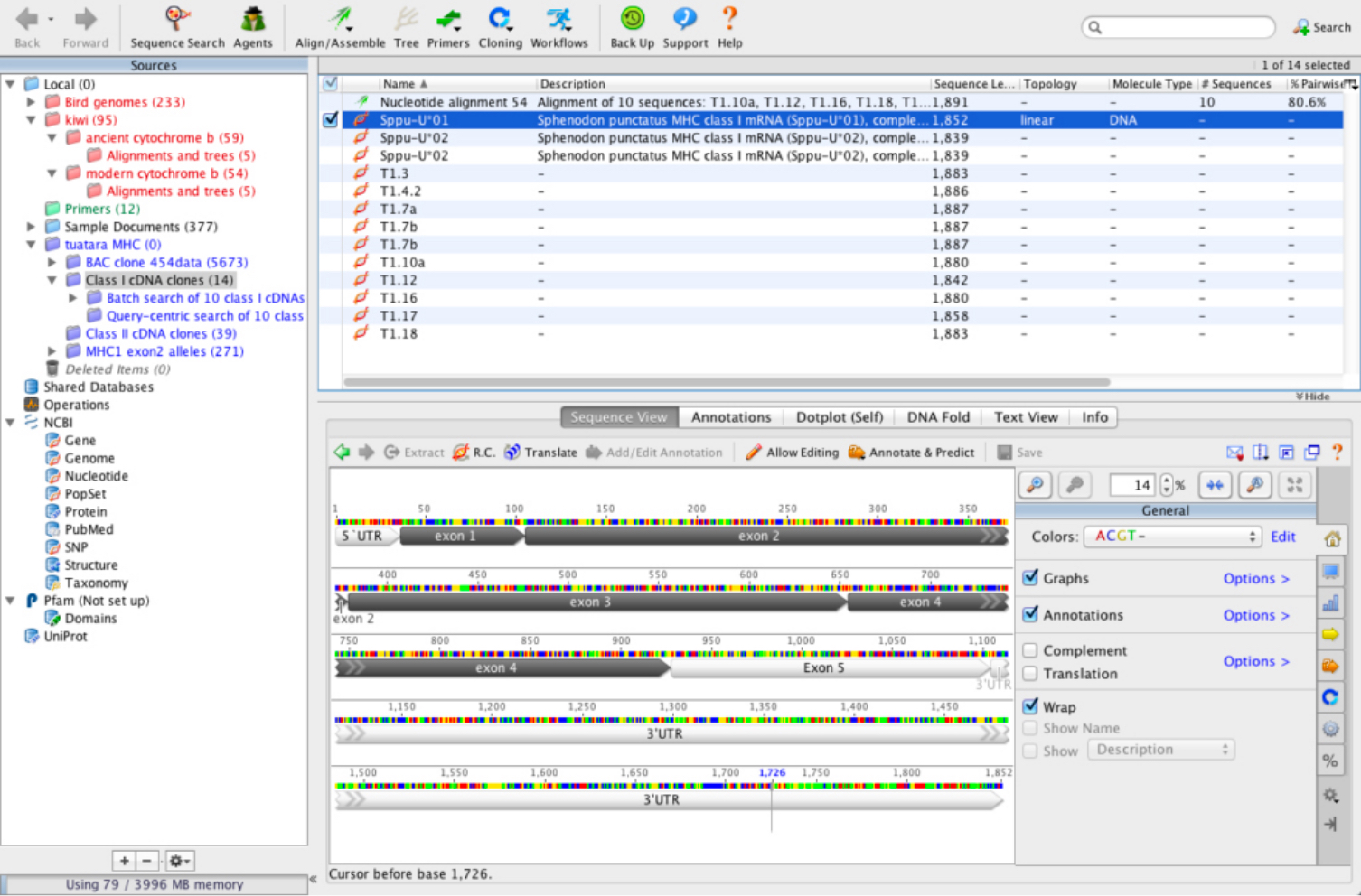
If there is a fasta file with the same name, and in the same folder as the annotation file, Geneious will ask if you wish to import this file as well.Ģ. When you import the annotation file you will be prompted for the sequence in one of two ways:ġ. 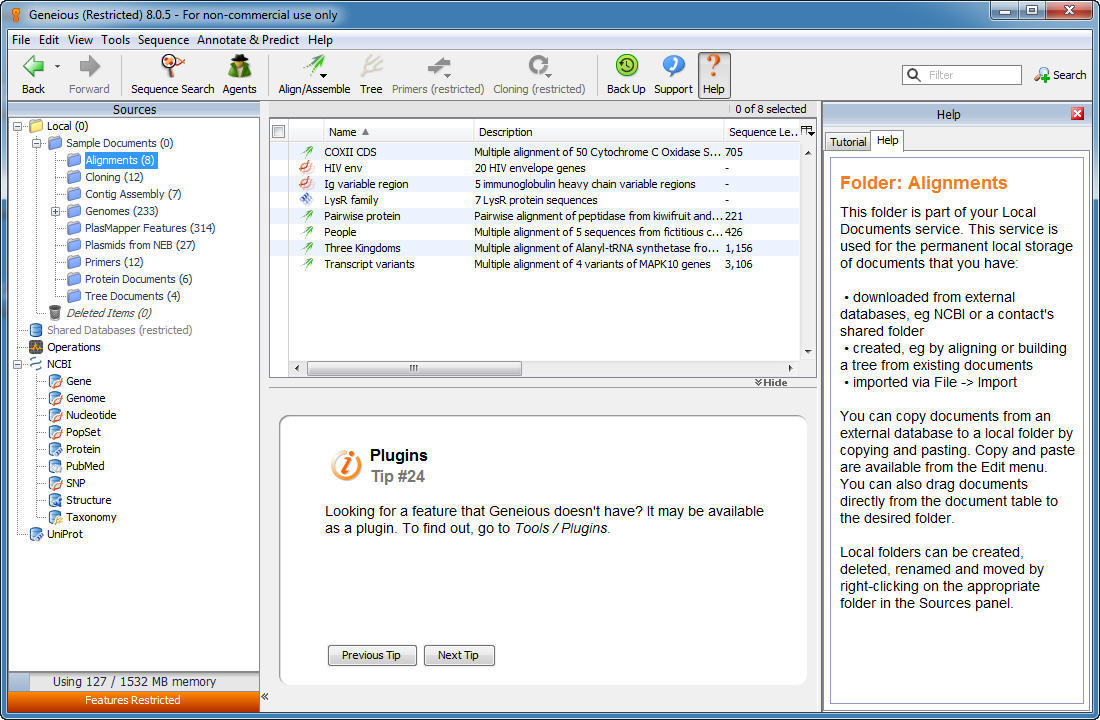
To do this, you can either import the sequence from a fasta file at the same time you import the annotation file, or you can import the file onto an existing sequence in your Geneious database. As these files generally do not contain sequence, you must provide the sequence to import the annotations on to. GFF and BED files normally contain gene and other sequence features, while VCF files are used for variant call data. GFF, BED and VCF are commonly used annotation file formats.
0 Comments
Leave a Reply. |
 RSS Feed
RSS Feed
